Project information
- Project date: March 1, 2026- April 10, 2026
- Tools: pyMTL RTLsim, Synopsys VCS (rtl, ffgl, baglsim), Synopsys PT (sta, pwr), Cadence Innovus (PNR), Mentor Calibre (LVS, DRC)
- Link to Presentation
Summary
DNA Local Sequence Alignment Accelerator:
ASIC Tapeout
Digital ASIC Design Project
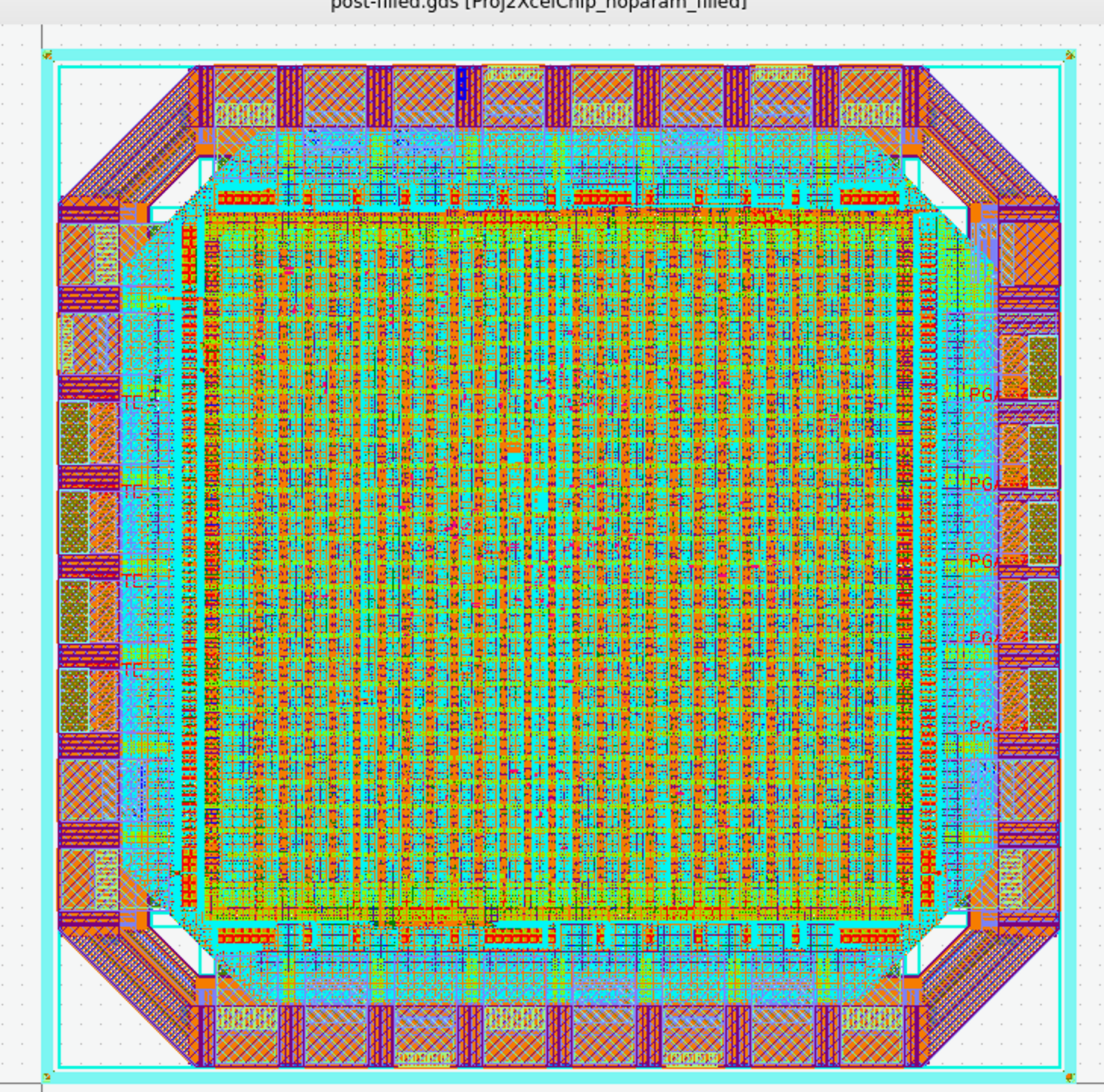
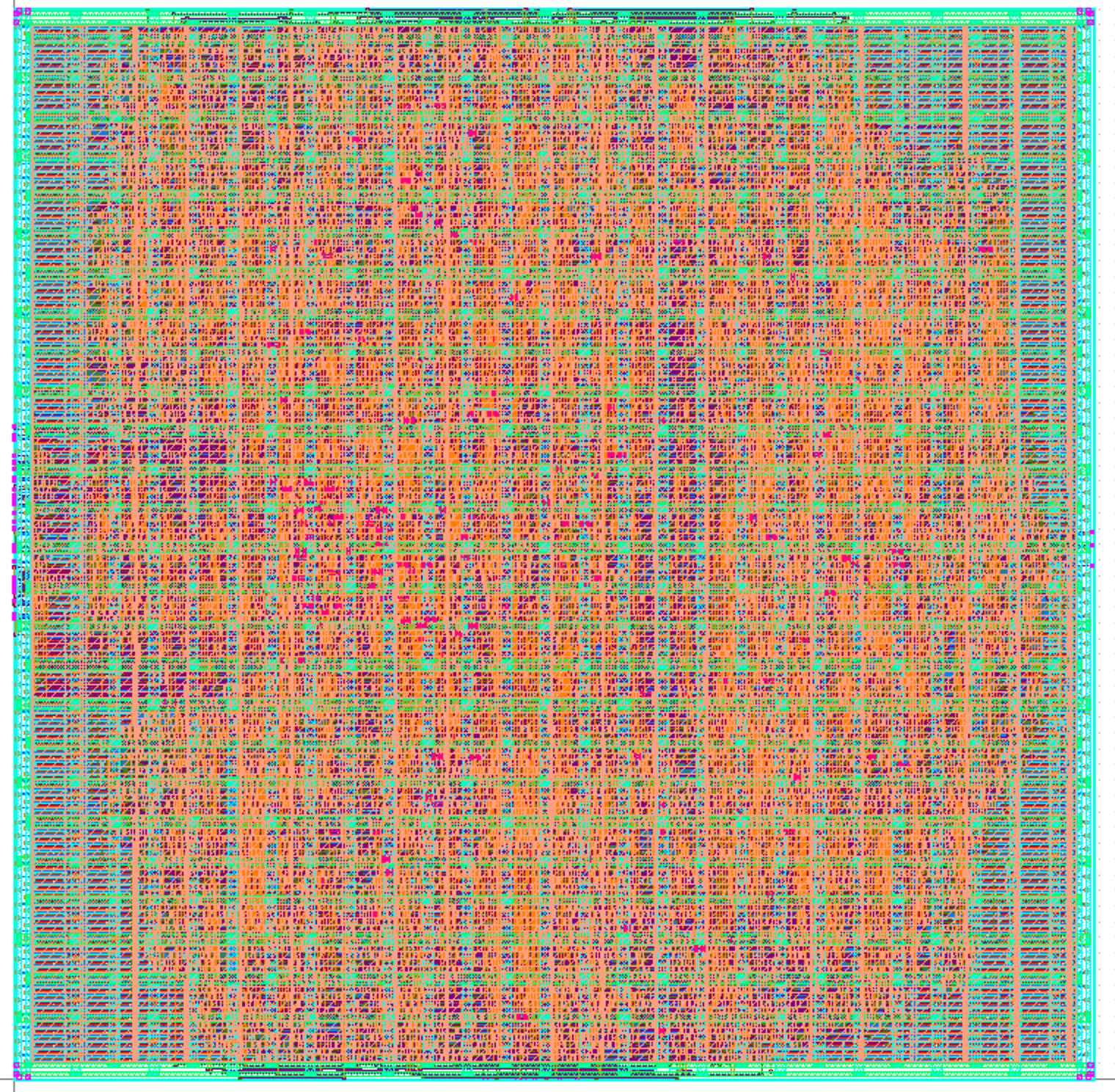
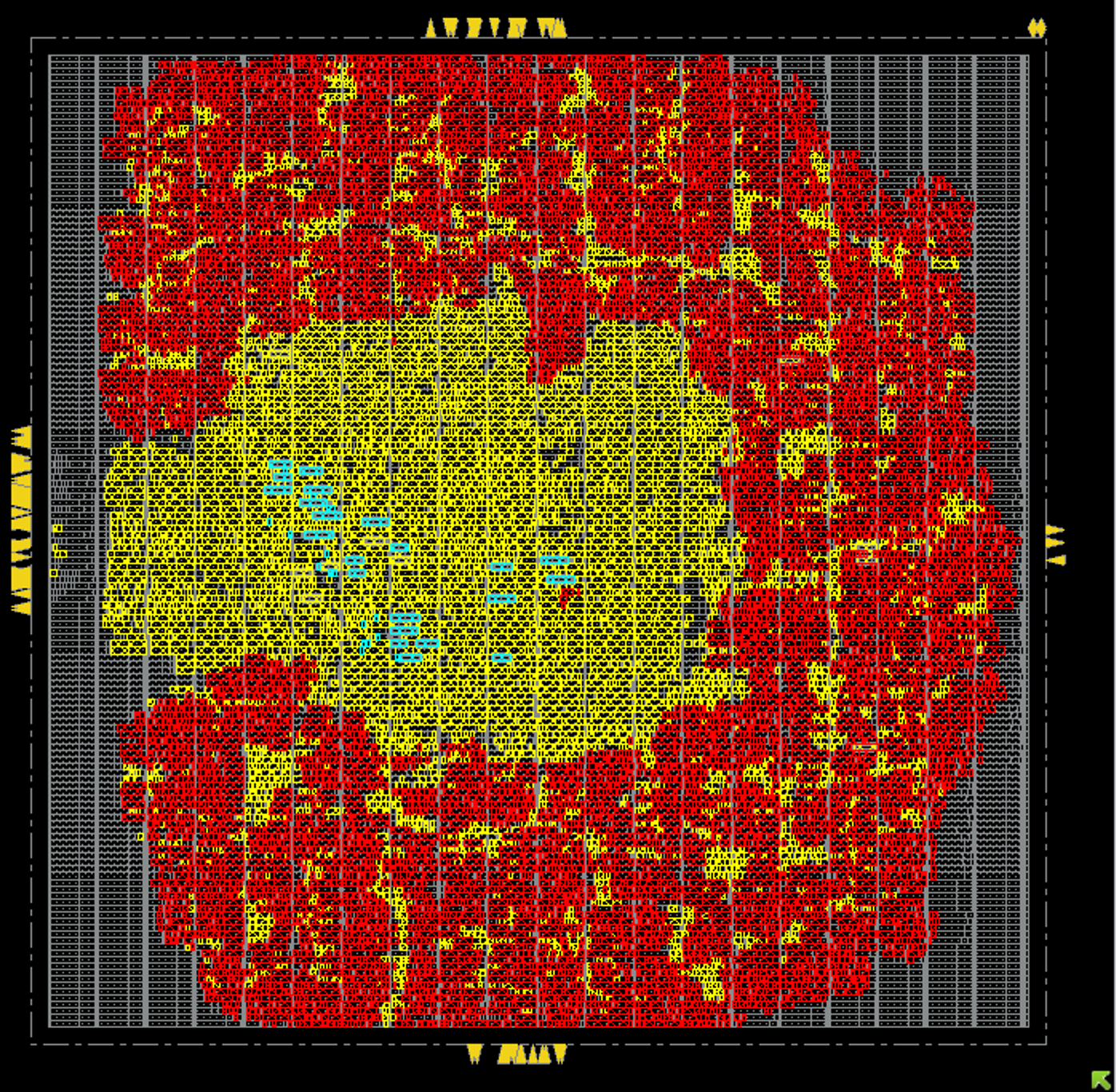
As part of Cornell’s Digital ASIC Design course (ECE 6745), my group and I designed a 1mmx1mm ASIC in the TSMC 180nm node for tape out this comming summer
The ASIC implements a systolic array of processing elements to accelerate the Smith-Waterman algorithm for local sequence alignment, which is widely used in bioinformatics for comparing DNA sequences. The design includes a custom controller for the processing elements, and we developed a custom RTL design, synthesis, place and route flow using industry-standard tools. We also performed extensive verification and testing to ensure the correctness and performance of the design.